Color Deconvolution
This article describes how to implement color deconvolution and un-mixing method for RGB images in arivis Vision4D.
See also
Introduction
Color or RGB images are common in histopathology where bright-field images capture white light from which the diagnostic dyes absorb a certain portion of the spectrum. These images can be segmented in a range of ways, but a better separation of the signal components from the red, green, and blue channels can significantly improve segmentation results. This script is an implementation in Vision4D of the method developed by Ruifrok, A.C. & Johnston, D.A. as described here.
This method has been developed specifically for the unmixing of diaminobenzidine, hematoxylin and eosin dies.
The output of the script is the creation of three additional channels to the current image set containing the unmixed signal.
This script relies on the NumPy libraries that are installed automatically with version 3.4 or above of arivis Vision4D. Earlier versions may require additional components.
Applying the script to an image
Once all the necessary edits have been made we can run the script on an image. If the script editor is still open, the script can be run by clicking on Script>Run Script or by pressing the F5 key on the keyboard.
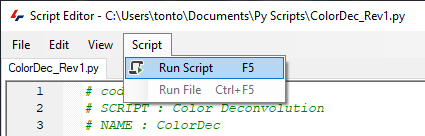
Note that the script will run on the active image set. If more sets need to be processed, you can select these additional sets in the viewer and run the script again or modifying the script to process multiple sets.
The script can also be run directly from the Vision4D window by going to Extras> Run Script... and selecting the .py file that has been saved after making the necessary changes described above.
Interpreting the results
The output of the script is three additional channels in the source image sets. These channels are named for the dye signal that has been extracted and can be used in any pipeline as any regular image channel for segmentation etc. In cases of dual staining (e.g. H&E), the third channel essentially contains only the signal that could not be reliably attached to either of the other two. this means that this third channel can essentially be used as some form of quality control for the output since in a perfect deconvolution this third channel would ideally be empty. If there is still a strong signal in the third channel this usually indicates that the deconvolution parameters do not perfectly match the staining in the image. This could be caused by white balancing issues or extreme staining (either too weak or too strong).
For more information on the process of the deconvolution, please refer to the published literature on which this is based, linked above.
Conclusions
Once the script has been run, pipelines can be created as per usual that can use the newly created channel for segmentation or any additional modification, just like any other image channel.
Support for this script can be obtained by logging a support ticket using the link at the top of this page.